Submission guidelines
Contents
-
Instructions for Authors
- Types of papers
- 1. General information
- Manuscript Submission
- Title page
- Text
- References
- Tables
- Artwork and Illustrations Guidelines
- Supplementary Information (SI)
- After acceptance
- Open Choice
- Research Data Policy and Data Availability Statements
- Scientific style
- Additional information for Scientific style
- Ethical Responsibilities of Authors
- Competing Interests
- Research involving human participants, their data or biological material
- Research involving animals, their data or biological material
- Informed consent
- Authorship principles
- Research involving human embryos, gametes and stem cells
- Utilization of plants, algae, fungi
- Editorial procedure
- Editing Services
- Open access publishing
- Mistakes to avoid during manuscript preparation
Instructions for Authors
Types of papers
- Comments
- Reviews
- Brief Reports
- Original Research Articles
1. General information
Medicinal Chemistry Research is a journal for the prompt disclosure of novel experimental achievements in many facets of drug design, drug discovery, and the elucidation of mechanisms of action of biologically active compounds. Articles are sought which emphasize research in chemical biological relationships, especially with respect to: structure-activity relationships, investigations of biochemical and pharmacological targets of drug action, and correlations of structures with the mode of action of biologically active compounds. Studies will be welcomed that increase our understanding of biochemical interactions between drug molecules, ions, free radicals, and sterically important sections of macromolecular targets. The Journal is also dedicated to medicinal plants and to bioactive natural products of plant, fungal, mammalian, and aquatic origin. The Journal publishes original contributions in the following major areas:
- Design, synthesis, and structure-activity relationships of bioactive compounds
- Docking, molecular modeling, and computational studies of bioactive interactions
- Characterization of active ingredients of medicinal plants and identification of bioactivity in plant extracts
- Identification of targets and mechanism of activity of bioactive compounds
- Chemistry and biochemistry of bioactive natural products of plant origin
- Critical reviews of the historical, clinical, and legal status of medicinal agents, and accounts on topical issues.
Contributions reporting the following are not normally considered for publication:
- Biological activity on crude extracts that have not been characterized by analysis of their secondary metabolites (HPLC, 1H and 13C NMR including 2D NMR).
- Unexceptional and predictable bioactivity (e.g. antioxidant properties of phenolic or antibacterial activity of essential oils or antioxidant properties of metals such as iron, copper, etc.).
- Uncritical ethnopharmacological investigations, where a list of plants and their use are simply reported.
- Synthetic work in which the spectroscopic characterization is not complete (e.g., 1H and 13C NMR, HRMS, CHN, UV, IR, etc.).
- Computational work that simply discusses the docking, molecular modeling, QSAR, SAR, and computational studies of bioactive interactions without validation of the method (with experimental data).
- Biological activity that is low and insufficient to generate meaningful structure activity relationship.
Violation of any of the following rules will result in an immediate rejection:
RULE 1: The manuscript does not fall into any of the areas of interest of the Journal.
RULE 2: The manuscript is too preliminary (e.g., data without comparison to a reference, or without a positive control).
RULE 3: The botanical source is not clearly identified, authenticated, or documented (voucher specimen).
RULE 4: The manuscript is too focused on a non-chemical subject (e.g., pharmacology, analytical studies of active ingredients, analytical studies of drug concentrations (ADME is suitable), etc.
RULE 5: Manuscripts that simply discuss antioxidant properties of phenols or other compounds known to possess antioxidant effects.
RULE 6: Computer QSAR/modeling manuscripts that lack experimental biological validation of the proposed model(s).
RULE 7: The manuscript does not follow the formatting provided in this document.
RULE 8: The manuscripts contains poor English or difficult to read language.
2. General Considerations and types of manuscripts
Authors are strongly encouraged to provide their manuscript in an electronic format. The text must be in a single-column format and lines with double space. Use plain font 12 point Times New Roman and symbols (use internationally accepted signs and symbols for units, SI units). Use the automatic page numbering function to number all the pages. Ensure that all special characters are presented in the body of the text and do not use graphics. Abbreviations, except for very common ones, must be defined the first time they are used and a list supplied with the manuscript.
Using clear and concise English will help the editors and the reviewers concentrate on the scientific merit of the manuscript and thus smooth the peer review process. We reject manuscripts with good science that are poorly written.
The text of a research manuscript should be divided into the following sections: Introduction, Results and Discussion, Conclusions, Materials and Methods/Experimental, Acknowledgements (Funding), Conflict of Interest, and References as appropriate for the specific manuscript type. Tables, figures, and schemes should be embedded in the text or be included right after the references on separate pages (one each per page). Do not upload tables, figures, and schemes that are to be published in the manuscript into the electronic supplementary material. Authors are encouraged to provide supplementary material to keep the manuscript to a reasonable length appropriate for the specific manuscript type.
Types of manuscript accepted include comments, reviews, brief reports, and original research articles.
2.1. Guidelines for comments
Comments are normally by invitation only. Unsolicited comment articles are rarely considered, but if you wish to enquire further about the suitability of your article, you can email Dr. Hans Detlef Klueber (detlef.klueber@springer.com).
Comments are short focused articles or viewpoints that are usually invited by the journal. These articles are generally peer-reviewed in consultation with the journal's Editorial Board. A comment article can be editorial in nature discussing new topics or new technologies in medicinal chemistry research and can also be used to highlight one or more exciting research articles recently published in Med. Chem. Res.
Comments should be less than 3,000 words with less than 15 references, preferably <5 references. No abstract or graphical abstract is needed. Supplementary materials are allowed.
2.2. Guidelines for review articles
Review articles are peer-reviewed articles covering important topic areas that are of significant interest to the broad community of scientists working in drug design, discovery, and development. They are typically 10-20 pages in length. We also welcome shorter, mini-review style articles under this article type that cover new drugs recently developed and tested in clinical trials and/or received regulatory approval.
Review articles do not need to be comprehensive and should focus on important drug targets, mechanism of action, and/or important structure-activity relationship findings and key physicochemical/pharmaceutical/DMPK properties that made the molecule into a drug. Authors should summarize important findings rather than reiterating details published in the primary literature. Whenever possible, authors should make their own summary figures instead of reusing multiple figures from the primary literature. If reusing or simply adapting figures already published, authors need to obtain permission from the copy right holders and reference accordingly.
Authors should include balanced coverage of the field reviewed and not just their own research. The authors are encouraged to incorporate their own perspectives and do their own analysis of data published. In the final section, the authors should discuss future directions and trends where appropriate.
2.3. Guidelines for brief reports
Brief reports are peer-reviewed short papers that present original and significant results in drug design, discovery, and development for rapid dissemination or short notes that present technical developments and innovations in the use of new technologies such as artificial intelligence, machine learning, high throughput experimentation, flow chemistry, and software to advance drug discovery and development.
Brief reports are limited to 6 pages in length. The paper should contain an abstract, graphical abstract, main body and references, and contain no more than 6 figures, schemes, or tables, combined. The abstract is limited to 150 words. The paper may not need an experimental section but details about compound characterization and purity confirmation can be placed in a supporting document.
2.4. Guidelines for original research articles
Original research articles are peer-reviewed full papers that present original and significant results in drug design, discovery, and development with an experimental section. There are no page limits, although most original research articles would have around 8-15 journal pages.
The text of an original research article should be divided into the following sections: Introduction, Results and Discussion, Conclusions, Materials and Methods/Experimental, Acknowledgements (Funding), Conflict of Interest, and References. Tables, figures, and schemes should be embedded in the text or be included right after the references on separate pages (one each per page). The authors should limit the number of schemes/figures/tables to a total of 10 or less in the manuscript.
2.5. Manuscript requirements at a glance
| Article type | Brief description | Word count/ # print pages/ # illustrations | Abstract length | Graphical abstract | # Keywords | # references |
|---|---|---|---|---|---|---|
| 1. Comments | <3,000 words ≤6 print pages With or without figures +schemes +tables | NOT NEEDED | NOT NEEDED | Optional | ≤15, preferably <5 | |
| 2. Reviews |
| ~5,000 -10,000 words ~10-20 print pages ~10 figures +schemes +tables, combined | ≤250 words | YES | 4-6 | ~100-200 |
| 3. Brief Reports |
| <3,000 words ≤6 print pages ≤6 figures +schemes +tables, combined | ≤150 words | YES | 4-6 | ≤30 |
| 4. Original Research Articles |
| ~5,000 -10,000 words ~10-20 print pages ~10 figures +schemes +tables, combined | ≤250 words | YES | 4-6 | ~50 |
Graphical Abstract. The author(s) are strongly encouraged to provide a graphical abstract that is a single schematic image which visually represents the main findings of the article, allowing readers to easily capture the content of the article at a single glance. The graphical abstract must meet the same quality and permissions standards as any other figure in the article. A caption is not needed, while compound numbers can be given in the graphical abstract. The use of color is encouraged and there is no charge for the graphical abstract. The graphical abstract should be legible at a size of 5 × 3 inches (130 × 75 mm, w × h)) using a regular screen resolution of 96 dpi. Graphical abstracts can be selected as the front cover image.
Cover Image. Authors are encouraged to submit a high resolution image of their graphical abstract as a separate file for consideration as a cover image at the revision/proofing stage. The image file should be created at publication size: 8 × 4.8 in (200 × 120 mm, w × h) at a resolution of at least 300 dpi in uncompressed JPEG, TIFF, PNG, PSD, AI, EPS, or PDF format.
Immediately after the abstract paragraph/graphical abstract, 4 to 6 keywords, should be provided for indexing purposes under the heading Keywords.
Manuscript Submission
Manuscript Submission
Submission of a manuscript implies: that the work described has not been published before; that it is not under consideration for publication anywhere else; that its publication has been approved by all co-authors, if any, as well as by the responsible authorities – tacitly or explicitly – at the institute where the work has been carried out. The publisher will not be held legally responsible should there be any claims for compensation.
Permissions
Authors wishing to include figures, tables, or text passages that have already been published elsewhere are required to obtain permission from the copyright owner(s) for both the print and online format and to include evidence that such permission has been granted when submitting their papers. Any material received without such evidence will be assumed to originate from the authors.
Online Submission
Please follow the hyperlink “Submit manuscript” on the right and upload all of your manuscript files following the instructions given on the screen.
Source Files
Please ensure you provide all relevant editable source files at every submission and revision. Failing to submit a complete set of editable source files will result in your article not being considered for review. For your manuscript text please always submit in common word processing formats such as .docx or LaTeX.
Title page
Please make sure your title page contains the following information
Title
The title should be concise and informative.
Author information
- The name(s) of the author(s)
- The affiliation(s) of the author(s), i.e. institution, (department), city, (state), country
- A clear indication and an active e-mail address of the corresponding author
- If available, the 16-digit ORCID of the author(s)
If address information is provided with the affiliation(s) it will also be published.
For authors that are (temporarily) unaffiliated we will only capture their city and country of residence, not their e-mail address unless specifically requested.
Large Language Models (LLMs), such as ChatGPT, do not currently satisfy our authorship criteria. Notably an attribution of authorship carries with it accountability for the work, which cannot be effectively applied to LLMs. Use of an LLM should be properly documented in the Methods section (and if a Methods section is not available, in a suitable alternative part) of the manuscript.
Abstract
Please provide an abstract of 150 to 250 words. The abstract should not contain any undefined abbreviations or unspecified references.
For life science journals only (when applicable)
- Trial registration number and date of registration for prospectively registered trials
- Trial registration number and date of registration, followed by “retrospectively registered”, for retrospectively registered trials
Keywords
Please provide 4 to 6 keywords which can be used for indexing purposes.
Statements and Declarations
The following statements should be included under the heading "Statements and Declarations" for inclusion in the published paper. Please note that submissions that do not include relevant declarations will be returned as incomplete.
- Competing Interests: Authors are required to disclose financial or non-financial interests that are directly or indirectly related to the work submitted for publication. Please refer to "Competing Interests and Funding" below for more information on how to complete this section.
Please see the relevant sections in the submission guidelines for further information as well as various examples of wording. Please revise/customize the sample statements according to your own needs.
Text
Text Formatting
Manuscripts should be submitted in Word.
- Use a normal, plain font (e.g., 10-point Times Roman) for text.
- Use italics for emphasis.
- Use the automatic page numbering function to number the pages.
- Do not use field functions.
- Use tab stops or other commands for indents, not the space bar.
- Use the table function, not spreadsheets, to make tables.
- Use the equation editor or MathType for equations.
- Save your file in docx format (Word 2007 or higher) or doc format (older Word versions).
Manuscripts with mathematical content can also be submitted in LaTeX. We recommend using Springer Nature’s LaTeX template.
Headings
Please use no more than three levels of displayed headings.
Abbreviations
Abbreviations should be defined at first mention and used consistently thereafter.
Footnotes
Footnotes can be used to give additional information, which may include the citation of a reference included in the reference list. They should not consist solely of a reference citation, and they should never include the bibliographic details of a reference. They should also not contain any figures or tables.
Footnotes to the text are numbered consecutively; those to tables should be indicated by superscript lower-case letters (or asterisks for significance values and other statistical data). Footnotes to the title or the authors of the article are not given reference symbols.
Always use footnotes instead of endnotes.
Acknowledgments
Acknowledgments of people, grants, funds, etc. should be placed in a separate section after Experimental. The names of funding organizations should be written in full.
References
Citation
Reference citations in the text should be identified by numbers in square brackets. Some examples:
1. Negotiation research spans many disciplines [3].
2. This result was later contradicted by Becker and Seligman [5].
3. This effect has been widely studied [1-3, 7].
Reference list
The list of references should only include works that are cited in the text and that have been published or accepted for publication. Personal communications and unpublished works should only be mentioned in the text.
The entries in the list should be numbered consecutively.
If available, please always include DOIs as full DOI links in your reference list (e.g. “https://doi.org/abc”).
- Journal article
Smith JJ. The world of science. Am J Sci. 1999;36:234–5.
- Article by DOI
Slifka MK, Whitton JL. Clinical implications of dysregulated cytokine production. J Mol Med. 2000; https://doi.org/10.1007/s001090000086
- Book
Blenkinsopp A, Paxton P. Symptoms in the pharmacy: a guide to the management of common illness. 3rd ed. Oxford: Blackwell Science; 1998.
- Book chapter
Wyllie AH, Kerr JFR, Currie AR. Cell death: the significance of apoptosis. In: Bourne GH, Danielli JF, Jeon KW, editors. International review of cytology. London: Academic; 1980. pp. 251–306.
- Online document
Doe J. Title of subordinate document. In: The dictionary of substances and their effects. Royal Society of Chemistry. 1999. http://www.rsc.org/dose/title of subordinate document. Accessed 15 Jan 1999.
Always use the standard abbreviation of a journal’s name according to the ISSN List of Title Word Abbreviations, see
If you are unsure, please use the full journal title.
Endnote
For authors using EndNote, Springer provides an output style that supports the formatting of in-text citations and reference list. Please click link below to download the EndNote style file.
Tables
- All tables are to be numbered using Arabic numerals.
- Tables should always be cited in text in consecutive numerical order.
- For each table, please supply a table caption (title) explaining the components of the table.
- Identify any previously published material by giving the original source in the form of a reference at the end of the table caption.
- Footnotes to tables should be indicated by superscript lower-case letters (or asterisks for significance values and other statistical data) and included beneath the table body.
Artwork and Illustrations Guidelines
Electronic Figure Submission
- Supply all figures electronically.
- Indicate what graphics program was used to create the artwork.
- For vector graphics, the preferred format is EPS; for halftones, please use TIFF format. MSOffice files are also acceptable.
- Vector graphics containing fonts must have the fonts embedded in the files.
- Name your figure files with "Fig" and the figure number, e.g., Fig1.eps.
Line Art
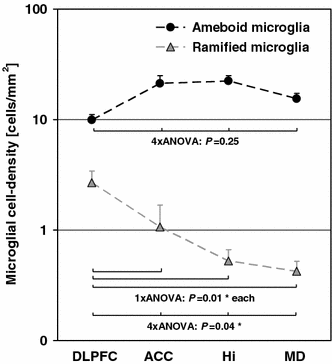
- Definition: Black and white graphic with no shading.
- Do not use faint lines and/or lettering and check that all lines and lettering within the figures are legible at final size.
- All lines should be at least 0.1 mm (0.3 pt) wide.
- Scanned line drawings and line drawings in bitmap format should have a minimum resolution of 1200 dpi.
- Vector graphics containing fonts must have the fonts embedded in the files.
Halftone Art
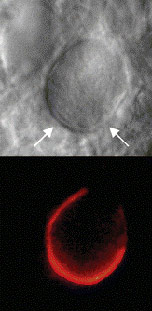
- Definition: Photographs, drawings, or paintings with fine shading, etc.
- If any magnification is used in the photographs, indicate this by using scale bars within the figures themselves.
- Halftones should have a minimum resolution of 300 dpi.
Combination Art

- Definition: a combination of halftone and line art, e.g., halftones containing line drawing, extensive lettering, color diagrams, etc.
- Combination artwork should have a minimum resolution of 600 dpi.
Color Art
- Color art is free of charge for online publication.
- If black and white will be shown in the print version, make sure that the main information will still be visible. Many colors are not distinguishable from one another when converted to black and white. A simple way to check this is to make a xerographic copy to see if the necessary distinctions between the different colors are still apparent.
- If the figures will be printed in black and white, do not refer to color in the captions.
- Color illustrations should be submitted as RGB (8 bits per channel).
Figure Lettering
- To add lettering, it is best to use Helvetica or Arial (sans serif fonts).
- Keep lettering consistently sized throughout your final-sized artwork, usually about 2–3 mm (8–12 pt).
- Variance of type size within an illustration should be minimal, e.g., do not use 8-pt type on an axis and 20-pt type for the axis label.
- Avoid effects such as shading, outline letters, etc.
- Do not include titles or captions within your illustrations.
Figure Numbering
- All figures are to be numbered using Arabic numerals.
- Figures should always be cited in text in consecutive numerical order.
- Figure parts should be denoted by lowercase letters (a, b, c, etc.).
- If an appendix appears in your article and it contains one or more figures, continue the consecutive numbering of the main text. Do not number the appendix figures,"A1, A2, A3, etc." Figures in online appendices [Supplementary Information (SI)] should, however, be numbered separately.
Figure Captions
- Each figure should have a concise caption describing accurately what the figure depicts. Include the captions in the text file of the manuscript, not in the figure file.
- Figure captions begin with the term Fig. in bold type, followed by the figure number, also in bold type.
- No punctuation is to be included after the number, nor is any punctuation to be placed at the end of the caption.
- Identify all elements found in the figure in the figure caption; and use boxes, circles, etc., as coordinate points in graphs.
- Identify previously published material by giving the original source in the form of a reference citation at the end of the figure caption.
Figure Placement and Size
- Figures should be submitted separately from the text, if possible.
- When preparing your figures, size figures to fit in the column width.
- For large-sized journals the figures should be 84 mm (for double-column text areas), or 174 mm (for single-column text areas) wide and not higher than 234 mm.
- For small-sized journals, the figures should be 119 mm wide and not higher than 195 mm.
Permissions
If you include figures that have already been published elsewhere, you must obtain permission from the copyright owner(s) for both the print and online format. Please be aware that some publishers do not grant electronic rights for free and that Springer will not be able to refund any costs that may have occurred to receive these permissions. In such cases, material from other sources should be used.
Accessibility
In order to give people of all abilities and disabilities access to the content of your figures, please make sure that
- All figures have descriptive captions (blind users could then use a text-to-speech software or a text-to-Braille hardware)
- Patterns are used instead of or in addition to colors for conveying information (colorblind users would then be able to distinguish the visual elements)
- Any figure lettering has a contrast ratio of at least 4.5:1
Generative AI Images
Please check Springer’s policy on generative AI images and make sure your work adheres to the principles described therein.
Supplementary Information (SI)
Springer accepts electronic multimedia files (animations, movies, audio, etc.) and other supplementary files to be published online along with an article or a book chapter. This feature can add dimension to the author's article, as certain information cannot be printed or is more convenient in electronic form.
Before submitting research datasets as Supplementary Information, authors should read the journal’s Research data policy. We encourage research data to be archived in data repositories wherever possible.
Submission
- Supply all supplementary material in standard file formats.
- Please include in each file the following information: article title, journal name, author names; affiliation and e-mail address of the corresponding author.
- To accommodate user downloads, please keep in mind that larger-sized files may require very long download times and that some users may experience other problems during downloading.
- High resolution (streamable quality) videos can be submitted up to a maximum of 25GB; low resolution videos should not be larger than 5GB.
Audio, Video, and Animations
- Aspect ratio: 16:9 or 4:3
- Maximum file size: 25 GB for high resolution files; 5 GB for low resolution files
- Minimum video duration: 1 sec
- Supported file formats: avi, wmv, mp4, mov, m2p, mp2, mpg, mpeg, flv, mxf, mts, m4v, 3gp
Text and Presentations
- Submit your material in PDF format; .doc or .ppt files are not suitable for long-term viability.
- A collection of figures may also be combined in a PDF file.
Spreadsheets
- Spreadsheets should be submitted as .csv or .xlsx files (MS Excel).
Specialized Formats
- Specialized formats such as .pdb (chemical), .wrl (VRML), .nb (Mathematica notebook), and .tex can also be supplied.
Collecting Multiple Files
- It is possible to collect multiple files in a .zip or .gz file.
Numbering
- If supplying any supplementary material, the text must make specific mention of the material as a citation, similar to that of figures and tables.
- Refer to the supplementary files as “Online Resource”, e.g., "... as shown in the animation (Online Resource 3)", “... additional data are given in Online Resource 4”.
- Name the files consecutively, e.g. “ESM_3.mpg”, “ESM_4.pdf”.
Captions
- For each supplementary material, please supply a concise caption describing the content of the file.
Processing of supplementary files
- Supplementary Information (SI) will be published as received from the author without any conversion, editing, or reformatting.
Accessibility
In order to give people of all abilities and disabilities access to the content of your supplementary files, please make sure that
- The manuscript contains a descriptive caption for each supplementary material
- Video files do not contain anything that flashes more than three times per second (so that users prone to seizures caused by such effects are not put at risk)
Generative AI Images
Please check Springer’s policy on generative AI images and make sure your work adheres to the principles described therein.
After acceptance
Upon acceptance, your article will be exported to Production to undergo typesetting. Shortly after this you will receive two e-mails. One contains a request to confirm your affiliation, choose the publishing model for your article, as well as to arrange rights and payment of any associated publication cost. A second e-mail containing a link to your article’s proofs will be sent once typesetting is completed.
Article publishing agreement
Depending on the ownership of the journal and its policies, you will either grant the Publisher an exclusive licence to publish the article or will be asked to transfer copyright of the article to the Publisher.
Offprints
Offprints can be ordered by the corresponding author.
Color illustrations
Online publication of color illustrations is free of charge. For color in the print version, authors will be expected to make a contribution towards the extra costs.
Proof reading
The purpose of the proof is to check for typesetting or conversion errors and the completeness and accuracy of the text, tables and figures. Substantial changes in content, e.g., new results, corrected values, title and authorship, are not allowed without the approval of the Editor.
After online publication, further changes can only be made in the form of an Erratum, which will be hyperlinked to the article.
Online First
The article will be published online after receipt of the corrected proofs. This is the official first publication citable with the DOI. After release of the printed version, the paper can also be cited by issue and page numbers.
Open Choice
Open Choice allows you to publish open access in more than 1850 Springer Nature journals, making your research more visible and accessible immediately on publication.
Article processing charges (APCs) vary by journal – view the full list
Benefits:
- Increased researcher engagement: Open Choice enables access by anyone with an internet connection, immediately on publication.
- Higher visibility and impact: In Springer hybrid journals, OA articles are accessed 4 times more often on average, and cited 1.7 more times on average*.
- Easy compliance with funder and institutional mandates: Many funders require open access publishing, and some take compliance into account when assessing future grant applications.
It is easy to find funding to support open access – please see our funding and support pages for more information.
*) Within the first three years of publication. Springer Nature hybrid journal OA impact analysis, 2018.
Copyright and license term – CC BY
Open Choice articles do not require transfer of copyright as the copyright remains with the author. In opting for open access, the author(s) agree to publish the article under the Creative Commons Attribution License.
Research Data Policy and Data Availability Statements
This journal operates a type 2 research data policy (life sciences).
The journal strongly encourages that all datasets on which the conclusions of the paper rely should be available to readers. We encourage authors to ensure that their datasets are either deposited in publicly available repositories (where available and appropriate) or presented in the main manuscript or additional supporting files whenever possible. Please see Springer Nature’s information on recommended repositories.
General repositories - for all types of research data - such as figshare and Dryad may be used where appropriate.
Datasets that are assigned digital object identifiers (DOIs) by a data repository may be cited in the reference list. Data citations should include the minimum information recommended by DataCite: authors, title, publisher (repository name), identifier.
Where a widely established research community expectation for data archiving in public repositories exists, submission to a community-endorsed, public repository is mandatory. Persistent identifiers (such as DOIs and accession numbers) for relevant datasets must be provided in the paper.
If the journal that you’re submitting to uses double-blind peer review and you are providing reviewers with access to your data (for example via a repository link, supplementary information or data on request), it is strongly suggested that the authorship in the data is also blinded. There are data repositories that can assist with this and/or will create a link to mask the authorship of your data.
For the following types of data set, submission to a community-endorsed, public repository is mandatory:
| Mandatory deposition | Suitable repositories |
| Protein sequences | Uniprot |
| DNA and RNA sequences | Genbank DNA DataBank of Japan (DDBJ) EMBL Nucleotide Sequence Database (ENA) |
| DNA and RNA sequencing data | NCBI Trace Archive NCBI Sequence Read Archive (SRA) |
| Genetic polymorphisms | dbSNP dbVar European Variation Archive (EVA) |
| Linked genotype and phenotype data | dbGAP The European Genome-phenome Archive (EGA) |
| Macromolecular structure | Worldwide Protein Data Bank (wwPDB) Biological Magnetic Resonance Data Bank (BMRB) Electron Microscopy Data Bank (EMDB) |
| Microarray data (must be MIAME compliant) | Gene Expression Omnibus (GEO) ArrayExpress |
| Crystallographic data for small molecules | Cambridge Structural Database |
For more information:
Research Data Policy Frequently Asked Questions
Data availability
The journal encourages authors to provide a statement of Data availability in their article. Data availability statements should include information on where data supporting the results reported in the article can be found, including, where applicable, hyperlinks to publicly archived datasets analysed or generated during the study. Data availability statements can also indicate whether data are available on request from the authors and where no data are available, if appropriate.
Data Availability statements can take one of the following forms (or a combination of more than one if required for multiple datasets):
- 1. The datasets generated during and/or analysed during the current study are available in the [NAME] repository, [PERSISTENT WEB LINK TO DATASETS]
- 2. The datasets generated during and/or analysed during the current study are not publicly available due [REASON WHY DATA ARE NOT PUBLIC] but are available from the corresponding author on reasonable request.
- 3. The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
- 4. Data sharing not applicable to this article as no datasets were generated or analysed during the current study.
- 5. All data generated or analysed during this study are included in this published article [and its supplementary information files].
More examples of template data availability statements, which include examples of openly available and restricted access datasets, are available:
Authors who need help understanding our data sharing policies, help finding a suitable data repository, or help organising and sharing research data can access our Author Support portal for additional guidance.
Scientific style
- Please always use internationally accepted signs and symbols for units (SI units).
- Nomenclature: Insofar as possible, authors should use systematic names similar to those used by IUPAC.
- Genus and species names should be in italics.
- Generic names of drugs and pesticides are preferred; if trade names are used, the generic name should be given at first mention.
- Please use the standard mathematical notation for formulae, symbols, etc.: Italic for single letters that denote mathematical constants, variables, and unknown quantities; Roman/upright for numerals, operators, and punctuation, and commonly defined functions or abbreviations, e.g., cos, det, e or exp, lim, log, max, min, sin, tan, d (for derivative); Bold for vectors, tensors, and matrices.
Additional information for Scientific style
Chemical Abstract Service (https://www.cas.org/) or IUPAC (https://iupac.org/)
- Manuscripts submitted to the journal are expected to adhere to internationally accepted nomenclature for receptors (http://www.guidetopharmacology.org/) and enzymes (https://www.enzyme-database.org/).
Ethical Responsibilities of Authors
This journal is committed to upholding the integrity of the scientific record. As a member of the Committee on Publication Ethics (COPE) the journal will follow the COPE guidelines on how to deal with potential acts of misconduct.
Authors should refrain from misrepresenting research results which could damage the trust in the journal, the professionalism of scientific authorship, and ultimately the entire scientific endeavour. Maintaining integrity of the research and its presentation is helped by following the rules of good scientific practice, which include*:
- The manuscript should not be submitted to more than one journal for simultaneous consideration.
- The submitted work should be original and should not have been published elsewhere in any form or language (partially or in full), unless the new work concerns an expansion of previous work. (Please provide transparency on the re-use of material to avoid the concerns about text-recycling (‘self-plagiarism’).
- A single study should not be split up into several parts to increase the quantity of submissions and submitted to various journals or to one journal over time (i.e. ‘salami-slicing/publishing’).
- Concurrent or secondary publication is sometimes justifiable, provided certain conditions are met. Examples include: translations or a manuscript that is intended for a different group of readers.
- Results should be presented clearly, honestly, and without fabrication, falsification or inappropriate data manipulation (including image based manipulation). Authors should adhere to discipline-specific rules for acquiring, selecting and processing data.
- No data, text, or theories by others are presented as if they were the author’s own (‘plagiarism’). Proper acknowledgements to other works must be given (this includes material that is closely copied (near verbatim), summarized and/or paraphrased), quotation marks (to indicate words taken from another source) are used for verbatim copying of material, and permissions secured for material that is copyrighted.
Important note: the journal may use software to screen for plagiarism.
- Authors should make sure they have permissions for the use of software, questionnaires/(web) surveys and scales in their studies (if appropriate).
- Research articles and non-research articles (e.g. Opinion, Review, and Commentary articles) must cite appropriate and relevant literature in support of the claims made. Excessive and inappropriate self-citation or coordinated efforts among several authors to collectively self-cite is strongly discouraged.
- Authors should avoid untrue statements about an entity (who can be an individual person or a company) or descriptions of their behavior or actions that could potentially be seen as personal attacks or allegations about that person.
- Research that may be misapplied to pose a threat to public health or national security should be clearly identified in the manuscript (e.g. dual use of research). Examples include creation of harmful consequences of biological agents or toxins, disruption of immunity of vaccines, unusual hazards in the use of chemicals, weaponization of research/technology (amongst others).
- Authors are strongly advised to ensure the author group, the Corresponding Author, and the order of authors are all correct at submission. Adding and/or deleting authors during the revision stages is generally not permitted, but in some cases may be warranted. Reasons for changes in authorship should be explained in detail. Please note that changes to authorship cannot be made after acceptance of a manuscript.
*All of the above are guidelines and authors need to make sure to respect third parties rights such as copyright and/or moral rights.
Upon request authors should be prepared to send relevant documentation or data in order to verify the validity of the results presented. This could be in the form of raw data, samples, records, etc. Sensitive information in the form of confidential or proprietary data is excluded.
If there is suspicion of misbehavior or alleged fraud the Journal and/or Publisher will carry out an investigation following COPE guidelines. If, after investigation, there are valid concerns, the author(s) concerned will be contacted under their given e-mail address and given an opportunity to address the issue. Depending on the situation, this may result in the Journal’s and/or Publisher’s implementation of the following measures, including, but not limited to:
- If the manuscript is still under consideration, it may be rejected and returned to the author.
- If the article has already been published online, depending on the nature and severity of the infraction:
- an erratum/correction may be placed with the article
- an expression of concern may be placed with the article
- or in severe cases retraction of the article may occur.
The reason will be given in the published erratum/correction, expression of concern or retraction note. Please note that retraction means that the article is maintained on the platform, watermarked “retracted” and the explanation for the retraction is provided in a note linked to the watermarked article.
- The author’s institution may be informed
- A notice of suspected transgression of ethical standards in the peer review system may be included as part of the author’s and article’s bibliographic record.
Fundamental errors
Authors have an obligation to correct mistakes once they discover a significant error or inaccuracy in their published article. The author(s) is/are requested to contact the journal and explain in what sense the error is impacting the article. A decision on how to correct the literature will depend on the nature of the error. This may be a correction or retraction. The retraction note should provide transparency which parts of the article are impacted by the error.
Suggesting / excluding reviewers
Authors are welcome to suggest suitable reviewers and/or request the exclusion of certain individuals when they submit their manuscripts. When suggesting reviewers, authors should make sure they are totally independent and not connected to the work in any way. It is strongly recommended to suggest a mix of reviewers from different countries and different institutions. When suggesting reviewers, the Corresponding Author must provide an institutional email address for each suggested reviewer, or, if this is not possible to include other means of verifying the identity such as a link to a personal homepage, a link to the publication record or a researcher or author ID in the submission letter. Please note that the Journal may not use the suggestions, but suggestions are appreciated and may help facilitate the peer review process.
Competing Interests
Authors are requested to disclose interests that are directly or indirectly related to the work submitted for publication. Interests within the last 3 years of beginning the work (conducting the research and preparing the work for submission) should be reported. Interests outside the 3-year time frame must be disclosed if they could reasonably be perceived as influencing the submitted work. Disclosure of interests provides a complete and transparent process and helps readers form their own judgments of potential bias. This is not meant to imply that a financial relationship with an organization that sponsored the research or compensation received for consultancy work is inappropriate.
Editorial Board Members and Editors are required to declare any competing interests and may be excluded from the peer review process if a competing interest exists. In addition, they should exclude themselves from handling manuscripts in cases where there is a competing interest. This may include – but is not limited to – having previously published with one or more of the authors, and sharing the same institution as one or more of the authors. Where an Editor or Editorial Board Member is on the author list we recommend they declare this in the competing interests section on the submitted manuscript. If they are an author or have any other competing interest regarding a specific manuscript, another Editor or member of the Editorial Board will be assigned to assume responsibility for overseeing peer review. These submissions are subject to the exact same review process as any other manuscript. Editorial Board Members are welcome to submit papers to the journal. These submissions are not given any priority over other manuscripts, and Editorial Board Member status has no bearing on editorial consideration.
Interests that should be considered and disclosed but are not limited to the following:
Funding: Research grants from funding agencies (please give the research funder and the grant number) and/or research support (including salaries, equipment, supplies, reimbursement for attending symposia, and other expenses) by organizations that may gain or lose financially through publication of this manuscript.
Employment: Recent (while engaged in the research project), present or anticipated employment by any organization that may gain or lose financially through publication of this manuscript. This includes multiple affiliations (if applicable).
Financial interests: Stocks or shares in companies (including holdings of spouse and/or children) that may gain or lose financially through publication of this manuscript; consultation fees or other forms of remuneration from organizations that may gain or lose financially; patents or patent applications whose value may be affected by publication of this manuscript.
It is difficult to specify a threshold at which a financial interest becomes significant, any such figure is necessarily arbitrary, so one possible practical guideline is the following: "Any undeclared financial interest that could embarrass the author were it to become publicly known after the work was published."
Non-financial interests: In addition, authors are requested to disclose interests that go beyond financial interests that could impart bias on the work submitted for publication such as professional interests, personal relationships or personal beliefs (amongst others). Examples include, but are not limited to: position on editorial board, advisory board or board of directors or other type of management relationships; writing and/or consulting for educational purposes; expert witness; mentoring relations; and so forth.
Primary research articles require a disclosure statement. Review articles present an expert synthesis of evidence and may be treated as an authoritative work on a subject. Review articles therefore require a disclosure statement. Other article types such as editorials, book reviews, comments (amongst others) may, dependent on their content, require a disclosure statement. If you are unclear whether your article type requires a disclosure statement, please contact the Editor-in-Chief.
Please note that, in addition to the above requirements, funding information (given that funding is a potential competing interest (as mentioned above)) needs to be disclosed upon submission of the manuscript in the peer review system. This information will automatically be added to the Record of CrossMark, however it is not added to the manuscript itself. Under ‘summary of requirements’ (see below) funding information should be included in the ‘Declarations’ section.
Summary of requirements
The above should be summarized in a statement and placed in a ‘Declarations’ section before the reference list under a heading of ‘Funding’ and/or ‘Competing interests’. Other declarations include Ethics approval, Consent, Data, Material and/or Code availability and Authors’ contribution statements.
Please see the various examples of wording below and revise/customize the sample statements according to your own needs.
When all authors have the same (or no) conflicts and/or funding it is sufficient to use one blanket statement.
Examples of statements to be used when funding has been received:
- Partial financial support was received from [...]
- The research leading to these results received funding from […] under Grant Agreement No[…].
- This study was funded by […]
- This work was supported by […] (Grant numbers […] and […]
Examples of statements to be used when there is no funding:
- The authors did not receive support from any organization for the submitted work.
- No funding was received to assist with the preparation of this manuscript.
- No funding was received for conducting this study.
- No funds, grants, or other support was received.
Examples of statements to be used when there are interests to declare:
- Financial interests: Author A has received research support from Company A. Author B has received a speaker honorarium from Company W and owns stock in Company X. Author C is consultant to company Y.
Non-financial interests: Author C is an unpaid member of committee Z.
- Financial interests: The authors declare they have no financial interests.
Non-financial interests: Author A is on the board of directors of Y and receives no compensation as member of the board of directors.
- Financial interests: Author A received a speaking fee from Y for Z. Author B receives a salary from association X. X where s/he is the Executive Director.
Non-financial interests: none.
- Financial interests: Author A and B declare they have no financial interests. Author C has received speaker and consultant honoraria from Company M and Company N. Dr. C has received speaker honorarium and research funding from Company M and Company O. Author D has received travel support from Company O.
Non-financial interests: Author D has served on advisory boards for Company M, Company N and Company O.
Examples of statements to be used when authors have nothing to declare:
- The authors have no relevant financial or non-financial interests to disclose.
- The authors have no competing interests to declare that are relevant to the content of this article.
- All authors certify that they have no affiliations with or involvement in any organization or entity with any financial interest or non-financial interest in the subject matter or materials discussed in this manuscript.
- The authors have no financial or proprietary interests in any material discussed in this article.
Authors are responsible for correctness of the statements provided in the manuscript. See also Authorship Principles. The Editor-in-Chief reserves the right to reject submissions that do not meet the guidelines described in this section.
Research involving human participants, their data or biological material
Ethics approval
When reporting a study that involved human participants, their data or biological material, authors should include a statement that confirms that the study was approved (or granted exemption) by the appropriate institutional and/or national research ethics committee (including the name of the ethics committee) and certify that the study was performed in accordance with the ethical standards as laid down in the 1964 Declaration of Helsinki and its later amendments or comparable ethical standards. If doubt exists whether the research was conducted in accordance with the 1964 Helsinki Declaration or comparable standards, the authors must explain the reasons for their approach, and demonstrate that an independent ethics committee or institutional review board explicitly approved the doubtful aspects of the study. If a study was granted exemption from requiring ethics approval, this should also be detailed in the manuscript (including the reasons for the exemption).
Retrospective ethics approval
If a study has not been granted ethics committee approval prior to commencing, retrospective ethics approval usually cannot be obtained and it may not be possible to consider the manuscript for peer review. The decision on whether to proceed to peer review in such cases is at the Editor's discretion.
Ethics approval for retrospective studies
Although retrospective studies are conducted on already available data or biological material (for which formal consent may not be needed or is difficult to obtain) ethics approval may be required dependent on the law and the national ethical guidelines of a country. Authors should check with their institution to make sure they are complying with the specific requirements of their country.
Ethics approval for case studies
Case reports require ethics approval. Most institutions will have specific policies on this subject. Authors should check with their institution to make sure they are complying with the specific requirements of their institution and seek ethics approval where needed. Authors should be aware to secure informed consent from the individual (or parent or guardian if the participant is a minor or incapable) See also section on Informed Consent.
Cell lines
If human cells are used, authors must declare in the manuscript: what cell lines were used by describing the source of the cell line, including when and from where it was obtained, whether the cell line has recently been authenticated and by what method. If cells were bought from a life science company the following need to be given in the manuscript: name of company (that provided the cells), cell type, number of cell line, and batch of cells.
It is recommended that authors check the NCBI database for misidentification and contamination of human cell lines. This step will alert authors to possible problems with the cell line and may save considerable time and effort.
Further information is available from the International Cell Line Authentication Committee (ICLAC).
Authors should include a statement that confirms that an institutional or independent ethics committee (including the name of the ethics committee) approved the study and that informed consent was obtained from the donor or next of kin.
Research Resource Identifiers (RRID)
Research Resource Identifiers (RRID) are persistent unique identifiers (effectively similar to a DOI) for research resources. This journal encourages authors to adopt RRIDs when reporting key biological resources (antibodies, cell lines, model organisms and tools) in their manuscripts.
Examples:
Organism: Filip1tm1a(KOMP)Wtsi RRID:MMRRC_055641-UCD
Cell Line: RST307 cell line RRID:CVCL_C321
Antibody: Luciferase antibody DSHB Cat# LUC-3, RRID:AB_2722109
Plasmid: mRuby3 plasmid RRID:Addgene_104005
Software: ImageJ Version 1.2.4 RRID:SCR_003070
RRIDs are provided by the Resource Identification Portal. Many commonly used research resources already have designated RRIDs. The portal also provides authors links so that they can quickly register a new resource and obtain an RRID.
Clinical Trial Registration
The World Health Organization (WHO) definition of a clinical trial is "any research study that prospectively assigns human participants or groups of humans to one or more health-related interventions to evaluate the effects on health outcomes". The WHO defines health interventions as “A health intervention is an act performed for, with or on behalf of a person or population whose purpose is to assess, improve, maintain, promote or modify health, functioning or health conditions” and a health-related outcome is generally defined as a change in the health of a person or population as a result of an intervention.
To ensure the integrity of the reporting of patient-centered trials, authors must register prospective clinical trials (phase II to IV trials) in suitable publicly available repositories. For example www.clinicaltrials.gov or any of the primary registries that participate in the WHO International Clinical Trials Registry Platform.
The trial registration number (TRN) and date of registration should be included as the last line of the manuscript abstract.
For clinical trials that have not been registered prospectively, authors are encouraged to register retrospectively to ensure the complete publication of all results. The trial registration number (TRN), date of registration and the words 'retrospectively registered’ should be included as the last line of the manuscript abstract.
Standards of reporting
Springer Nature advocates complete and transparent reporting of biomedical and biological research and research with biological applications. Authors are recommended to adhere to the minimum reporting guidelines hosted by the EQUATOR Network when preparing their manuscript.
Exact requirements may vary depending on the journal; please refer to the journal’s Instructions for Authors.
Checklists are available for a number of study designs, including:
Randomised trials (CONSORT) and Study protocols (SPIRIT)
Observational studies (STROBE)
Systematic reviews and meta-analyses (PRISMA) and protocols (Prisma-P)
Diagnostic/prognostic studies (STARD) and (TRIPOD)
Case reports (CARE)
Clinical practice guidelines (AGREE) and (RIGHT)
Qualitative research (SRQR) and (COREQ)
Animal pre-clinical studies (ARRIVE)
Quality improvement studies (SQUIRE)
Economic evaluations (CHEERS)
Summary of requirements
The above should be summarized in a statement and placed in a ‘Declarations’ section before the reference list under a heading of ‘Ethics approval’.
Please see the various examples of wording below and revise/customize the sample statements according to your own needs.
Examples of statements to be used when ethics approval has been obtained:
• All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. The study was approved by the Bioethics Committee of the Medical University of A (No. ...).
• This study was performed in line with the principles of the Declaration of Helsinki. Approval was granted by the Ethics Committee of University B (Date.../No. ...).
• Approval was obtained from the ethics committee of University C. The procedures used in this study adhere to the tenets of the Declaration of Helsinki.
• The questionnaire and methodology for this study was approved by the Human Research Ethics committee of the University of D (Ethics approval number: ...).
Examples of statements to be used for a retrospective study:
• Ethical approval was waived by the local Ethics Committee of University A in view of the retrospective nature of the study and all the procedures being performed were part of the routine care.
• This research study was conducted retrospectively from data obtained for clinical purposes. We consulted extensively with the IRB of XYZ who determined that our study did not need ethical approval. An IRB official waiver of ethical approval was granted from the IRB of XYZ.
• This retrospective chart review study involving human participants was in accordance with the ethical standards of the institutional and national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. The Human Investigation Committee (IRB) of University B approved this study.
Examples of statements to be used when no ethical approval is required/exemption granted:
• This is an observational study. The XYZ Research Ethics Committee has confirmed that no ethical approval is required.
• The data reproduced from Article X utilized human tissue that was procured via our Biobank AB, which provides de-identified samples. This study was reviewed and deemed exempt by our XYZ Institutional Review Board. The BioBank protocols are in accordance with the ethical standards of our institution and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Authors are responsible for correctness of the statements provided in the manuscript. See also Authorship Principles. The Editor-in-Chief reserves the right to reject submissions that do not meet the guidelines described in this section.
Research involving animals, their data or biological material
The welfare of animals (vertebrate and higher invertebrate) used for research, education and testing must be respected. Authors should supply detailed information on the ethical treatment of their animals in their submission. For that purpose they may use the ARRIVE checklist which is designed to be used when submitting manuscripts describing animal research.
For studies involving client-owned animals, authors must also document informed consent from the client or owner and adherence to a high standard (best practice) of veterinary care.
Authors are recommended to comply with:
• The International Union for Conservation of Nature (IUCN) Policy Statement on Research Involving Species at Risk of Extinction and consult the IUCN red list index of threatened species.
• Convention on the Trade in Endangered Species of Wild Fauna and Flora
When reporting results authors should indicate:
• … that the studies have been approved by a research ethics committee at the institution or practice at which the studies were conducted. Please provide the name of ethics committee and relevant permit number;
• … whether the legal requirements or guidelines in the country and/or state or province for the care and use of animals have been followed.
Researchers from countries without any legal requirements or guidelines voluntarily should refer to the following sites for guidance:
– The Basel Declaration describes fundamental principles of using animals in biomedical research
– The International Council for Laboratory Animal Science (ICLAS) provides ethical guidelines for researchers as well as editors and reviewers
– The Association for the study of Animal Behaviour describes ethical guidelines for the treatment of animals in research and teaching
– The International Association of Veterinary Editors’ Consensus Author Guidelines on Animal Ethics provide guidelines for authors on animal ethics and welfare
Researchers may wish to consult the most recent (ethical) guidelines available from relevant taxon-oriented professional societies.
If a study was granted exemption or did not require ethics approval, this should also be detailed in the manuscript.
Summary of requirements
The above should be summarized in a statement and placed in a ‘Declarations’ section before the reference list under a heading of ‘Ethics approval’.
Please see the various examples of wording below and revise/customize the sample statements according to your own needs.
Examples of statements to be used when ethics approval has been obtained:
• All procedures involving animals were in compliance with the European Community Council Directive of 24 November 1986, and ethical approval was granted by the Kocaeli University Ethics Committee (No. 29 12 2014, Kocaeli, Turkey).
• All procedures performed in the study were in accordance with the ARVO Statement for Use of Animals in Ophthalmic Vision and Research. The ethical principles established by the National Institutes of Health Guide for the Care and Use of Laboratory Animals (NIH Publications No. 8523, revised 2011) were followed. The research protocol was approved by the Ethics Committee on Animal Use (Protocol No. 06174/14) of FCAV/Unesp, Jaboticabal.
• This study involved a questionnaire-based survey of farmers as well as blood sampling from their animals. The study protocol was assessed and approved by Haramaya University, research and extension office. Participants provided their verbal informed consent for animal blood sampling as well as for the related survey questions. Collection of blood samples was carried out by veterinarians adhering to the regulations and guidelines on animal husbandry and welfare.
• All brown bear captures and handling were approved by the Ethical Committee on Animal Experiments, Uppsala, Sweden (Application C18/15) and the Swedish Environmental Protection Agency in compliance with Swedish laws and regulations.
• The ethics governing the use and conduct of experiments on animals were strictly observed, and the experimental protocol was approved by the University of Maiduguri Senate committee on Medical Research ethics. Proper permit and consent were obtained from the Maiduguri abattoir management, before the faecal samples of the cattle and camels slaughtered in this abattoir were used for this experiment.
Examples of statements to be used when no ethical approval is required/exemption granted:
• No approval of research ethics committees was required to accomplish the goals of this study because experimental work was conducted with an unregulated invertebrate species.
• As the trappings of small mammals were conducted as part of regular pest control measures in accordance with the NATO Standardized Agreement 2048 "Deployment Pest and Vector Surveillance and Control ", no approval by an ethics committee was required.
• All experiments have been conducted as per the guidelines of the Institutional Animal Ethics Committee, Department of Zoology, Utkal University, Bhubaneswar, Odisha, India. However, the insect species used in this study is reared for commercial production of raw silk materials, as a part of agro-based industry. Therefore, use of this animal in research does not require ethical clearance. We have obtained permission from the office of Research officer sericulture, Baripada, Orissa, India for the provision of infrastructure and support for rearing of silkworm both in indoor and outdoor conditions related to our study to promote sericulture practices.
Authors are responsible for correctness of the statements provided in the manuscript. See also Authorship Principles. The Editor-in-Chief reserves the right to reject submissions that do not meet the guidelines described in this section.
Informed consent
All individuals have individual rights that are not to be infringed. Individual participants in studies have, for example, the right to decide what happens to the (identifiable) personal data gathered, to what they have said during a study or an interview, as well as to any photograph that was taken. This is especially true concerning images of vulnerable people (e.g. minors, patients, refugees, etc) or the use of images in sensitive contexts. In many instances authors will need to secure written consent before including images.
Identifying details (names, dates of birth, identity numbers, biometrical characteristics (such as facial features, fingerprint, writing style, voice pattern, DNA or other distinguishing characteristic) and other information) of the participants that were studied should not be published in written descriptions, photographs, and genetic profiles unless the information is essential for scholarly purposes and the participant (or parent/guardian if the participant is a minor or incapable or legal representative) gave written informed consent for publication. Complete anonymity is difficult to achieve in some cases. Detailed descriptions of individual participants, whether of their whole bodies or of body sections, may lead to disclosure of their identity. Under certain circumstances consent is not required as long as information is anonymized and the submission does not include images that may identify the person.
Informed consent for publication should be obtained if there is any doubt. For example, masking the eye region in photographs of participants is inadequate protection of anonymity. If identifying characteristics are altered to protect anonymity, such as in genetic profiles, authors should provide assurance that alterations do not distort meaning.
Exceptions where it is not necessary to obtain consent:
• Images such as x rays, laparoscopic images, ultrasound images, brain scans, pathology slides unless there is a concern about identifying information in which case, authors should ensure that consent is obtained.
• Reuse of images: If images are being reused from prior publications, the Publisher will assume that the prior publication obtained the relevant information regarding consent. Authors should provide the appropriate attribution for republished images.
Consent and already available data and/or biologic material
Regardless of whether material is collected from living or dead patients, they (family or guardian if the deceased has not made a pre-mortem decision) must have given prior written consent. The aspect of confidentiality as well as any wishes from the deceased should be respected.
Data protection, confidentiality and privacy
When biological material is donated for or data is generated as part of a research project authors should ensure, as part of the informed consent procedure, that the participants are made aware what kind of (personal) data will be processed, how it will be used and for what purpose. In case of data acquired via a biobank/biorepository, it is possible they apply a broad consent which allows research participants to consent to a broad range of uses of their data and samples which is regarded by research ethics committees as specific enough to be considered “informed”. However, authors should always check the specific biobank/biorepository policies or any other type of data provider policies (in case of non-bio research) to be sure that this is the case.
Consent to Participate
For all research involving human subjects, freely-given, informed consent to participate in the study must be obtained from participants (or their parent or legal guardian in the case of children under 16) and a statement to this effect should appear in the manuscript. In the case of articles describing human transplantation studies, authors must include a statement declaring that no organs/tissues were obtained from prisoners and must also name the institution(s)/clinic(s)/department(s) via which organs/tissues were obtained. For manuscripts reporting studies involving vulnerable groups where there is the potential for coercion or where consent may not have been fully informed, extra care will be taken by the editor and may be referred to the Springer Nature Research Integrity Group.
Consent to Publish
Individuals may consent to participate in a study, but object to having their data published in a journal article. Authors should make sure to also seek consent from individuals to publish their data prior to submitting their paper to a journal. This is in particular applicable to case studies. A consent to publish form can be found
Summary of requirements
The above should be summarized in a statement and placed in a ‘Declarations’ section before the reference list under a heading of ‘Consent to participate’ and/or ‘Consent to publish’. Other declarations include Funding, Competing interests, Ethics approval, Consent, Data and/or Code availability and Authors’ contribution statements.
Please see the various examples of wording below and revise/customize the sample statements according to your own needs.
Sample statements for "Consent to participate":
Informed consent was obtained from all individual participants included in the study.
Informed consent was obtained from legal guardians.
Written informed consent was obtained from the parents.
Verbal informed consent was obtained prior to the interview.
Sample statements for “Consent to publish”:
The authors affirm that human research participants provided informed consent for publication of the images in Figure(s) 1a, 1b and 1c.
The participant has consented to the submission of the case report to the journal.
Patients signed informed consent regarding publishing their data and photographs.
Sample statements if identifying information about participants is available in the article:
Additional informed consent was obtained from all individual participants for whom identifying information is included in this article.
Authors are responsible for correctness of the statements provided in the manuscript. See also Authorship Principles. The Editor-in-Chief reserves the right to reject submissions that do not meet the guidelines described in this section.
Images will be removed from publication if authors have not obtained informed consent or the paper may be removed and replaced with a notice explaining the reason for removal.
Authorship principles
These guidelines describe authorship principles and good authorship practices to which prospective authors should adhere to.
Authorship clarified
The Journal and Publisher assume all authors agreed with the content and that all gave explicit consent to submit and that they obtained consent from the responsible authorities at the institute/organization where the work has been carried out, before the work is submitted.
The Publisher does not prescribe the kinds of contributions that warrant authorship. It is recommended that authors adhere to the guidelines for authorship that are applicable in their specific research field. In absence of specific guidelines it is recommended to adhere to the following guidelines*:
All authors whose names appear on the submission
1) made substantial contributions to the conception or design of the work; or the acquisition, analysis, or interpretation of data; or the creation of new software used in the work;
2) drafted the work or revised it critically for important intellectual content;
3) approved the version to be published; and
4) agree to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
* Based on/adapted from:
ICMJE, Defining the Role of Authors and Contributors,
Disclosures and declarations
All authors are requested to include information regarding sources of funding, financial or non-financial interests, study-specific approval by the appropriate ethics committee for research involving humans and/or animals, informed consent if the research involved human participants, and a statement on welfare of animals if the research involved animals (as appropriate).
The decision whether such information should be included is not only dependent on the scope of the journal, but also the scope of the article. Work submitted for publication may have implications for public health or general welfare and in those cases it is the responsibility of all authors to include the appropriate disclosures and declarations.
Data transparency
All authors are requested to make sure that all data and materials as well as software application or custom code support their published claims and comply with field standards. Please note that journals may have individual policies on (sharing) research data in concordance with disciplinary norms and expectations.
Role of the Corresponding Author
One author is assigned as Corresponding Author and acts on behalf of all co-authors and ensures that questions related to the accuracy or integrity of any part of the work are appropriately addressed.
The Corresponding Author is responsible for the following requirements:
- ensuring that all listed authors have approved the manuscript before submission, including the names and order of authors;
- managing all communication between the Journal and all co-authors, before and after publication;*
- providing transparency on re-use of material and mention any unpublished material (for example manuscripts in press) included in the manuscript in a cover letter to the Editor;
- making sure disclosures, declarations and transparency on data statements from all authors are included in the manuscript as appropriate (see above).
* The requirement of managing all communication between the journal and all co-authors during submission and proofing may be delegated to a Contact or Submitting Author. In this case please make sure the Corresponding Author is clearly indicated in the manuscript.
Author contributions
In absence of specific instructions and in research fields where it is possible to describe discrete efforts, the Publisher recommends authors to include contribution statements in the work that specifies the contribution of every author in order to promote transparency. These contributions should be listed at the separate title page.
Examples of such statement(s) are shown below:
• Free text:
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by [full name], [full name] and [full name]. The first draft of the manuscript was written by [full name] and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
• Conceptualization: [full name], …; Methodology: [full name], …; Formal analysis and investigation: [full name], …; Writing - original draft preparation: [full name, …]; Writing - review and editing: [full name], …; Funding acquisition: [full name], …; Resources: [full name], …; Supervision: [full name],….
For review articles where discrete statements are less applicable a statement should be included who had the idea for the article, who performed the literature search and data analysis, and who drafted and/or critically revised the work.
For articles that are based primarily on the student’s dissertation or thesis, it is recommended that the student is usually listed as principal author:
Affiliation
The primary affiliation for each author should be the institution where the majority of their work was done. If an author has subsequently moved, the current address may additionally be stated. Addresses will not be updated or changed after publication of the article.
Changes to authorship
Authors are strongly advised to ensure the correct author group, the Corresponding Author, and the order of authors at submission. Changes of authorship by adding or deleting authors, and/or changes in Corresponding Author, and/or changes in the sequence of authors are not accepted after acceptance of a manuscript.
- Please note that author names will be published exactly as they appear on the accepted submission!
Please make sure that the names of all authors are present and correctly spelled, and that addresses and affiliations are current.
Adding and/or deleting authors at revision stage are generally not permitted, but in some cases it may be warranted. Reasons for these changes in authorship should be explained. Approval of the change during revision is at the discretion of the Editor-in-Chief. Please note that journals may have individual policies on adding and/or deleting authors during revision stage.
Author identification
Authors are recommended to use their ORCID ID when submitting an article for consideration or acquire an ORCID ID via the submission process.
Deceased or incapacitated authors
For cases in which a co-author dies or is incapacitated during the writing, submission, or peer-review process, and the co-authors feel it is appropriate to include the author, co-authors should obtain approval from a (legal) representative which could be a direct relative.
Authorship issues or disputes
In the case of an authorship dispute during peer review or after acceptance and publication, the Journal will not be in a position to investigate or adjudicate. Authors will be asked to resolve the dispute themselves. If they are unable the Journal reserves the right to withdraw a manuscript from the editorial process or in case of a published paper raise the issue with the authors’ institution(s) and abide by its guidelines.
Confidentiality
Authors should treat all communication with the Journal as confidential which includes correspondence with direct representatives from the Journal such as Editors-in-Chief and/or Handling Editors and reviewers’ reports unless explicit consent has been received to share information.
Research involving human embryos, gametes and stem cells
Manuscripts that report experiments involving the use of human embryos and gametes, human embryonic stem cells and related materials, and clinical applications of stem cells must include confirmation that all experiments were performed in accordance with relevant guidelines and regulations (See also Research involving human participants, their data or biological material.
The manuscript should include an ethics statement identifying the institutional and/or national research ethics committee (including the name of the ethics committee) approving the experiments and describing any relevant details. Authors should confirm that informed consent (See also Informed Consent) was obtained from all recipients and/or donors of cells or tissues, where necessary, and describe the conditions of donation of materials for research, such as human embryos or gametes. Copies of approval and redacted consent documents may be requested by the Journal.
We encourage authors to follow the principles laid out in the ISSCR Guidelines for Stem Cell Research and Clinical Translation.
In deciding whether to publish papers describing modifications of the human germline, the Journal is guided by safety considerations, compliance with applicable regulations, as well as the status of the societal debate on the implications of such modifications for future generations. In case of concerns regarding a particular type of study the Journal may seek the advice from the Springer Nature Research Integrity Group.
The decision to publish a paper is the responsibility of the Editor-in-Chief of the Journal.
Utilization of plants, algae, fungi
This journal values stewardship, transparency, and adhering to governance with regards to collecting and utilizing specimens and conducting experiments and/or field studies. Therefore the journal sets out the following guidelines:
Field studies involving genetically engineered plants must be conducted in accordance with national or local legislation and, if applicable, the manuscript needs to include a statement specifying the appropriate permissions and/or licences.
Authors utilizing genetic plant resources received via local suppliers/collectors, such as species collected from protected areas or endangered species with medical importance, must conduct their experiments following the Nagoya Protocol (as part of the Convention on Biological Diversity).
Authors whose research is focusing on quarantine organisms (i.e. harmful or pest organisms, including plant pathogens) should adhere to national legislation and notify the relevant National Plant Protection Organization of new findings before publication. More information can be found via the International Plant Protection Convention.
In principle, it is recommended that authors comply with:
- The International Union for Conservation of Nature (IUCN) Policy Statement on Research Involving Species at Risk of Extinction and consult the IUCN red list index of threatened species
- Convention on the Trade in Endangered Species of Wild Fauna and Flora
Voucher specimens ensure that the identity of organisms studied in the field or in laboratory experiments can be verified, and ensure that new species concepts can be applied to past research. Voucher specimens documenting all investigated accessions (for population samples at least one specimen per population) are to be deposited in a public herbarium, for example: Index Herbariorum, or other public collection providing access to deposited material. Information on the voucher specimen and who identified it must be included in the manuscript such as Genus name, species name, author, and year of publication.
Names of plants, algae and fungi
Manuscripts containing new taxon names or other nomenclatural acts must follow the guidelines set by the International Code of Nomenclature for algae, fungi, and plants.
Authors describing new fungal taxa should register the names with a recognized repository, such as Mycobank, and request a unique digital identifier which should be included in the published article.
Editorial procedure
Single-blind peer review
This journal follows a single-blind reviewing procedure.
This journal also publishes special/guest-edited issues. The peer review process for these articles is the same as the peer review process of the journal in general.
Additionally, if a guest editor authors an article in their issue/collection, they will not handle the peer review process.
Additional Information
All submissions to Springer journals are first reviewed for completeness and only then sent to be assessed by an Editor who will decide whether they are suitable for peer review. Where an Editor is on the author list or has any other competing interest regarding a specific manuscript, another member of the Editorial Board will be assigned to oversee peer review. Editors will consider the peer-reviewed reports when making a decision, but are not bound by the opinions or recommendations therein. A concern raised by a single peer reviewer or the Editor themself may result in the manuscript being rejected. Authors receive peer review reports with the editorial decision on their manuscript.
This journal operates a single-blind peer-review system, where the reviewers are aware of the names and affiliations of the authors, but the reviewer reports provided to authors are anonymous.The benefit of single-blind peer review is that it is the traditional model of peer review that many reviewers are comfortable with, and it facilitates a dispassionate critique of a manuscript.
Submitted manuscripts will generally be reviewed by two or more experts who will be asked to evaluate whether the manuscript is scientifically sound and coherent, whether it duplicates already published work, and whether or not the manuscript is sufficiently clear for publication. The Editors will reach a decision based on these reports and, where necessary, they will consult with members of the Editorial Board.
When selecting reviewers, the editorial board seeks to avoid conflicts of interest. The journal accepts manuscript submissions from its own editorial board members in cases where the identities of the associate editor and referees handling the manuscript can remain fully confidential. To be accepted, manuscripts submitted by editorial board members must meet the same quality standards as all other accepted submissions.
Editing Services
English
How can you help improve your manuscript for publication?
Presenting your work in a well-structured manuscript and in well-written English gives it its best chance for editors and reviewers to understand it and evaluate it fairly. Many researchers find that getting some independent support helps them present their results in the best possible light. The experts at Springer Nature Author Services can help you with manuscript preparation—including English language editing, developmental comments, manuscript formatting, figure preparation, translation, and more.
You can also use our free Grammar Check tool for an evaluation of your work.
Please note that using these tools, or any other service, is not a requirement for publication, nor does it imply or guarantee that editors will accept the article, or even select it for peer review.
Chinese (中文)
您怎么做才有助于改进您的稿件以便顺利发表?
如果在结构精巧的稿件中用精心组织的英语展示您的作品,就能最大限度地让编辑和审稿人理解并公正评估您的作品。许多研究人员发现,获得一些独立支持有助于他们以尽可能美好的方式展示他们的成果。Springer Nature Author Services 的专家可帮助您准备稿件,具体包括润色英语表述、添加有见地的注释、为稿件排版、设计图表、翻译等。
您还可以使用我们的免费语法检查工具来评估您的作品。
请注意,使用这些工具或任何其他服务不是发表前必须满足的要求,也不暗示或保证相关文章定会被编辑接受(甚至未必会被选送同行评审)。
Japanese (日本語)
発表に備えて、論文を改善するにはどうすればよいでしょうか?
内容が適切に組み立てられ、質の高い英語で書かれた論文を投稿すれば、編集者や査読者が論文を理解し、公正に評価するための最善の機会となります。多くの研究者は、個別のサポートを受けることで、研究結果を可能な限り最高の形で発表できると思っています。Springer Nature Author Servicesのエキスパートが、英文の編集、建設的な提言、論文の書式、図の調整、翻訳など、論文の作成をサポートいたします。
原稿の評価に、無料の文法チェックツールもご利用いただけます。
これらのツールや他のサービスをご利用いただくことは、論文を掲載するための要件ではありません。また、編集者が論文を受理したり、査読に選定したりすることを示唆または保証するものではないことにご注意ください。
Korean (한국어)
게재를 위해 원고를 개선하려면 어떻게 해야 할까요?
여러분의 작품을 체계적인 원고로 발표하는 것은 편집자와 심사자가 여러분의 연구를 이해하고 공정하게 평가할 수 있는 최선의 기회를 제공합니다. 많은 연구자들은 어느 정도 독립적인 지원을 받는 것이 가능한 한 최선의 방법으로 자신의 결과를 발표하는 데 도움이 된다고 합니다. Springer Nature Author Services 전문가들은 영어 편집, 발전적인 논평, 원고 서식 지정, 그림 준비, 번역 등과 같은 원고 준비를 도와드릴 수 있습니다.
또한 당사의 무료 문법 검사도구를 사용하여 여러분의 연구를 평가할 수 있습니다.
이러한 도구 또는 기타 서비스를 사용하는 것은 게재를 위한 필수 요구사항이 아니며, 편집자가 해당 논문을 수락하거나 피어 리뷰에 해당 논문을 선택한다는 것을 암시하거나 보장하지는 않습니다.
Open access publishing
To find out more about publishing your work Open Access in Medicinal Chemistry Research, including information on fees, funding and licenses, visit our Open access publishing page.

