Overview
- Presents practical approaches to the pervasive question of how to choose parameter settings for sequence alignment
- Provides links to proven software implementations that work well on real data
- Introduces a general framework for parameter advising of broad utility in bioinformatics and beyond
Part of the book series: Computational Biology (COBO, volume 26)
Access this book
Tax calculation will be finalised at checkout
Other ways to access
About this book
This book develops a new approach called parameter advising for finding a parameter setting for a sequence aligner that yields a quality alignment of a given set of input sequences. In this framework, a parameter advisor is a procedure that automatically chooses a parameter setting for the input, and has two main ingredients:
(a) the set of parameter choices considered by the advisor, and
(b) an estimator of alignment accuracy used to rank alignments produced by the aligner.
On coupling a parameter advisor with an aligner, once the advisor is trained in a learning phase, the user simply inputs sequences to align, and receives an output alignment from the aligner, where the advisor has automatically selected the parameter setting.
The chapters first lay out the foundations of parameter advising, and then cover applications and extensions of advising. The content
• examines formulations of parameter advising and their computational complexity,
• develops methods for learning good accuracy estimators,
• presents approximation algorithms for finding good sets of parameter choices, and
• assesses software implementations of advising that perform well on real biological data.
Also explored are applications of parameter advising to
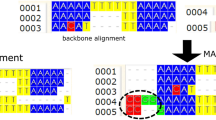
• adaptive local realignment, where advising is performed on local regions of the sequences to automatically adapt to varying mutation rates, and
• ensemble alignment, where advising is applied to an ensemble of aligners to effectively yield a new aligner of higher quality than the individual aligners in the ensemble.
The book concludes by offering future directions in advising research.
Similar content being viewed by others
Keywords
- Bioinformatics
- Biological sequence analysis
- Multiple sequence alignment
- Protein sequence alignment
- Alignment accuracy
- Alignment scoring parameters
- Substitution matrices
- Gap penalties
- Refining alignments
- Machine learning
- Ensemble methods
- Computational complexity
- Approximation algorithms
- Linear programming
- Integer linear programming
- algorithm analysis and problem complexity
Table of contents (10 chapters)
-
Front Matter
-
Foundations of Parameter Advising
-
Front Matter
-
-
Applications of Parameter Advising
-
Front Matter
-
-
Back Matter
Authors and Affiliations
Bibliographic Information
Book Title: Parameter Advising for Multiple Sequence Alignment
Authors: Dan DeBlasio, John Kececioglu
Series Title: Computational Biology
DOI: https://doi.org/10.1007/978-3-319-64918-4
Publisher: Springer Cham
eBook Packages: Computer Science, Computer Science (R0)
Copyright Information: Springer International Publishing AG 2017
Hardcover ISBN: 978-3-319-64917-7Published: 29 January 2018
Softcover ISBN: 978-3-319-87902-4Published: 06 June 2019
eBook ISBN: 978-3-319-64918-4Published: 04 January 2018
Series ISSN: 1568-2684
Series E-ISSN: 2662-2432
Edition Number: 1
Number of Pages: XIV, 152
Number of Illustrations: 2 b/w illustrations, 30 illustrations in colour
Topics: Computational Biology/Bioinformatics, Bioinformatics, Algorithm Analysis and Problem Complexity, Algorithms, Discrete Optimization